Publication Summary
Multiplexed Exchange-PAINT imaging reveals ligand-dependent EGFR and Met interactions in the plasma membrane
Jeffrey L. Werbin1,7, Maier S Avendaño2,3,7, Verena Becker1, Ralf Jungmann2,3,5, Peng Yin2,3, Gaudenz Danuser4,6, and Peter K. Sorger1
1HMS LINCS Center, Laboratory of Systems Pharmacology, Harvard Medical School, Boston, MA; 2 Department of Systems Biology, Harvard Medical School, Boston, MA; 3 Wyss Institute for Biologically Inspired Engineering, Harvard University, Boston, MA; 4 Department of Cell Biology, Harvard Medical School, Boston, MA; 5 Present address: Max Planck Institute of Biochemistry and LMU, Munich, Germany; 6 Present address: Department of Cell Biology, UT Southwestern Medical Center, Dallas, TX; 7 These authors contributed equally to this work.
Scientific Reports (2017) 7(1): 12150
doi:10.1038/s41598-017-12257-y / PMID: 28939861
Abstract
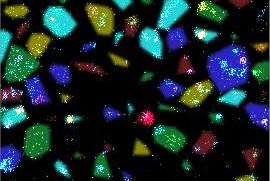
Signal transduction by receptor tyrosine kinases (RTKs) involves complex ligand- and time-dependent changes in conformation and modification state. High resolution structures are available for individual receptors dimers, but less is known about receptor clusters that form in plasma membranes composed of many different RTKs with the potential to interact. We report the use of multiplexed super-resolution imaging (Exchange-PAINT) followed by mean-shift clustering and random forest analysis to measure the precise distributions of five receptor tyrosine kinases (RTKs) from the ErbB, IGF-1R and Met families in breast cancer cells. We find that these receptors are intermixed nonhomogenously on the plasma membrane. Stimulation by EGF does not appear to induce a change in the density of EGFR in local clusters but instead results in formation of EGFR-Met and EGFR-ErbB3 associations; non-canonical EGFR-Met interactions are implicated in resistance to anti-cancer drugs but have not been previously detected in drug-naïve cells.
Available Data and Resources
| Data | Raw localization event data and mean-shift clustering results for EGFR, ErbB2, ErbB3, IGF1R and cMet receptors in serum-starved and EGF-stimulated BT20 triple-negative breast cancer cells. Note that this file is 3 GB in size and will take significant time to download. | Download (.zip) | |
| Code | MATLAB Source code for processing localization event data and performing mean-shift clustering. | Details | Download (.zip) |
Funding Sources
NIH grants CA112967-10 and U54 HL127365 to P.K.S; DP2OD007292, R01EB018659 and R21HD072481 to P.Y.; U01 GM067230 to G.D.; a postdoctoral fellowship from the American Cancer Society (PF-12-223-01-CCE) to J.L.W.; an HHMI International Student Research Fellowships to M.S.A.; an Alexander von Humboldt-Foundation through a Feodor-Lynen Fellowship to R.J.