Publication Summary
Growth rate inhibition metrics correct for confounders in measuring sensitivity to cancer drugs
Marc Hafner1,2, Mario Niepel1,2, Mirra Chung1, and Peter K. Sorger1
1HMS LINCS Center, Laboratory of Systems Pharmacology, Department of Systems Biology, Harvard Medical School, Boston, MA; 2MH and MN contributed equally to this work.
Nat Methods (2016) 13(6):521-527.
doi:10.1038/nmeth.3853 / PMID:27135972 / PMCID:PMC4887336
Synopsis
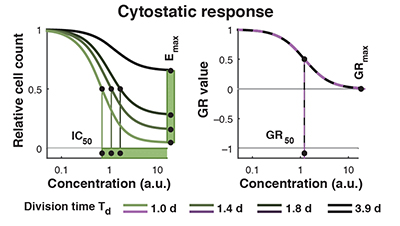
Conventional metrics for drug sensitivity and resistance such as IC50 or Emax are highly susceptible to artifacts arising from variation in cell division over the course of a response assay. We derive alternative growth rate inhibition (GR) metrics that are insensitive to division number and provide a more robust measure of the effects of perturbagens on dividing cells.
Key findings
- Traditional perturbagen-response metrics such as IC50, Emax, and AUC can vary dramatically, independent of the underlying biology, due to natural differences in cellular proliferation rate, variation in growth conditions, or changes in the duration of a dose-response experiment.
- Parameterization of growth rate data instead using normalized growth rate inhibition (GR) metrics yields GR50, GRmax, GRAOC, and hGR (Hill slope) values that are largely independent of cell division rate and assay duration.
- Only modest changes in experimental procedures are necessary to enable calculation of GR metrics from perturbagen-response datasets.
Abstract
Drug sensitivity and resistance are conventionally quantified by IC50 or Emax values, but these metrics are highly sensitive to the number of divisions taking place over the course of a response assay. The dependency of IC50 and Emax on division rate creates artifactual correlations between genotype and drug sensitivity, while obscuring valuable biological insights and interfering with biomarker discovery. We derive alternative small molecule drug-response metrics that are insensitive to division number. These are based on estimation of the magnitude of drug-induced growth rate inhibition (GR) using endpoint or time-course assays. We show that GR50 and GRmax are superior to conventional metrics for assessing the effects of small molecule drugs in dividing cells. Moreover, adopting GR metrics requires only modest changes in experimental protocols. We expect GR metrics to improve the study of cell signaling and growth using small molecules and biologics and to facilitate the discovery of drug-response biomarkers and the identification of drugs effective against specific patient-derived tumor cells.
Explore the method and associated data
We encourage readers to explore the mathematical basis of GR metrics and data arising from their application through the GR Browser, which presents a tutorial and interactive tool to help you learn more about GR metrics, an online GR Calculator to allow you to calculate and plot GR metrics from your own datasets, and visualization tools for dose-response datasets generated by the NIH LINCS Consortium. The GR Browser was developed through a collaboration between the HMS LINCS Center and the LINCS-BD2K Data Coordination and Integration Center. Matlab and Python scripts for offline calculation of GR metrics also are available through the Sorger lab GR50 Tools Git repository. Datasets described in Hafner, Niepel et al (2016) that assess the cell seeding density-dependence of GR metrics (Datasets #20256-20258) and that reanalyze dose-response data published in Heiser et al (2012)1 using GR metrics (Datasets #20268-20271) also are available via the HMS LINCS Database through the links below.
Available software
| Data | Cell counts and normalized GR values for each technical replicate of a breast cancer cell line study that assessed the effect of cell seeding density on drug dose response (HMS Dataset #20256) | Detail | Download (.xlsx) |
| Data | Mean cell counts and mean normalized GR values across technical replicates derived from HMS Dataset #20256 (HMS Dataset #20257) | Detail | Download (.xlsx) |
| Data | Calculated dose-response metrics derived from HMS Dataset #20257 (HMS Dataset #20258). | Detail | Download (.xlsx) |
| Data | Cell counts and normalized GR values across technical replicates for a panel of 71 breast cancer cell lines treated with a library of small molecule and antibody perturbagens (HMS Dataset #20268) | Detail | Download (.xlsx) |
| Data | Calculated dose-response metrics derived from HMS Dataset #20269 (HMS Dataset #20269) | Detail | Download (.xlsx) |
| Data | GR metric signatures derived from HMS Dataset #20269: Median, upper quartile, and lower quartile per perturbagen (HMS Dataset #20270) | Detail | Download (.xlsx) |
| Data | GR metric signatures derived from HMS Dataset #20268: Median, upper quartile, and lower quartile per perturbagen class (HMS Dataset #20271) | Detail | Download (.xlsx) |
| Software | The GR Browser: a suite of online tools for exploring and calculating GR metrics and visualizing the results of dose-response datasets | GR Browser | |
| Software | The GR50 Tools Git repository: public access to a GR metrics tutorial and the associated MATLAB scripts and Python code for calculating GR metrics from dose-response datasets | Git repository |
Funding sources
NIH LINCS grant U54-HL127365 and NIH grant P50-GM107618 to P.K.S. and a Swiss National Science Foundation fellowship (P300P3_147876) to M.H.
Related References
1. Heiser, L.M., Sadanandam, A., Kuo, W.L., Benz, S.C., Goldstein, T.C., Ng, S., Gibb, W.J., Wang, N.J., Ziyad, S., Tong, F., Bayani, N., Hu, Z., Billig, J.I., Dueregger, A., Lewis, S., Jakkula, L., Korkola, J.E., Durinck, S., Pepin, F., Guan, Y., Purdom, E., Neuvial, P., Bengtsson, H., Wood, K.W., Smith, P.G., Vassilev, L.T., Hennessy, B.T., Greshock, J., Bachman, K.E., Hardwicke, M.A., Park, J.W., Marton, L.J., Wolf, D.M., Collisson, E.A., Neve, R.M., Mills, G.B., Speed, T.P., Feiler, H.S., Wooster, R.F., Haussler, D., Stuart, J.M., Gray, J.W., and Spellman, P.T. (2012) Subtype and pathway specific responses to anticancer compounds in breast cancer. Proc Natl Acad Sci USA. 109(8):2724-9. doi:10.1073/pnas.1018854108 PMID:22003129 PMCID:PMC3286973